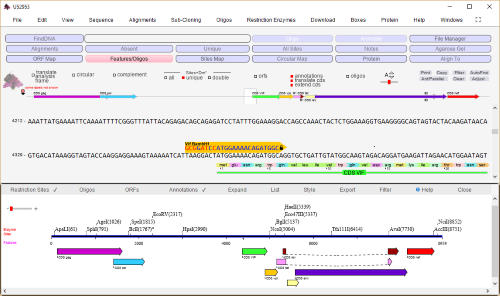
DNA Dynamo software features
sequence navigation and restriction site analysis
Genbank accession U52953
representing 'HIV-1 92BR025
from Brazil, complete genome'
was imported into
DNADynamo and a features
map displayed. Features may
be selected and/or set as the
ORF. Multi-partite (spliced)
features may be 'joined' and
opened in a new window. The
sequence display is set to
'translated mode' and is
illustrating the start of the 'vif'
gene, together with a selected
oligo.
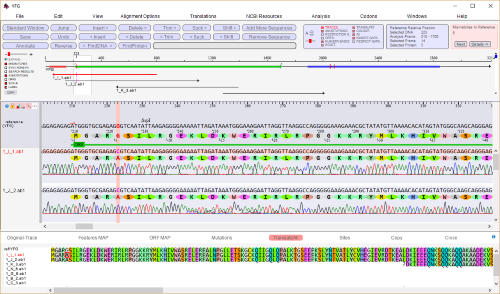
abi/scf chromatogram sequence alignment and editing
Editable sequence alignments in
the upper display area are linked
to chromatogram data in the lower
display area. A graphic map allows
easy visualization of sequencing
discrepancies while the 'guide'
sequence is translated and
illustrates annotations and oligos
selected from the 'features' map
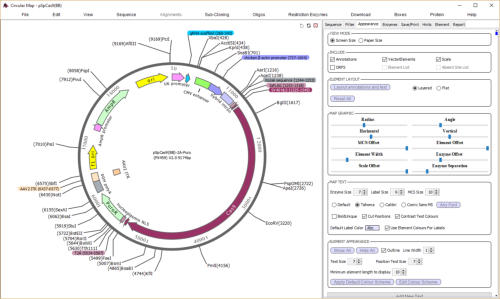
automatically add vector features and draw maps
DNADynamo’s circular maps are fully
editable. You can adjust the size,
colour and position of pretty much
everything, including setting your
favourite font, as well as adding
custom text labels. DNADynamo
maps also support a special MCS
display, and allow you to add a list of
features and/or a list of absent
restriction sites.
Additional Features
•
Oligo Editor and Database
•
Integrated web BLAST search and annotation
•
Annotated multiple sequence alignment via the DNADynamo ClustalW interface
•
Pattern Searching
•
Support for demo and licensed users - we answers all your questions as rapidly as we can
more information on DNA Dynamo software features and functions is available on the Getting Started with DNA Dynamo
page, and the Analysing Sequencing Data page,
DNA Dynamo software is available for OSX, Windows and Linux. Files created on one operating system are compatible
with other operating systems, making DNA Dynamo an ideal choice for mixed operating system environments.
Copyright © BlueTractorSoftware Ltd, all rights reserved

plan sub-cloning with fragment copy and replace, or
use the dedicated ‘Construct Maker
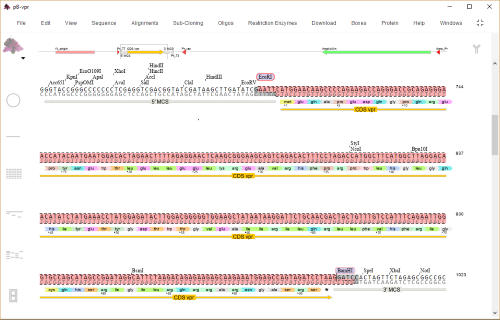
An EcoRI/BamHI fragment containing
the vpr gene is shown selected. The
fragment can be copied and then used
to replace a selected EcoRI/BamH1
fragment in another vector
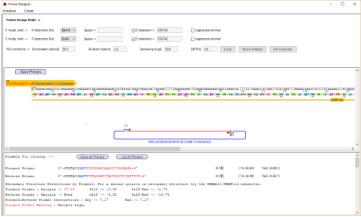
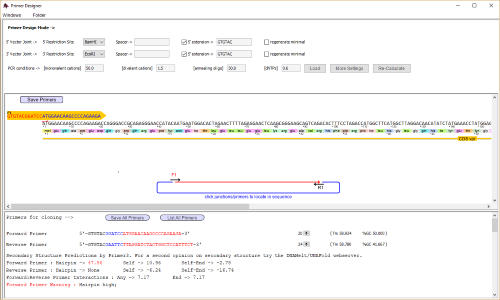
Design primers manually or use the primer designer to automatically
select primers with given properties eg for gibson/infusion cloning
or to add restriction sites
A fragment has been selected for use
in the Primer Designer - and primers
to add BamH1 sites and EcoRI sites
designed

DNADynamo
BlueTractorSoftware
•
Windows
•
OSX
•
Linux
DNA Dynamo software features
sequence navigation and restriction site analysis
Genbank accession U52953
representing 'HIV-1 92BR025
from Brazil, complete genome'
was imported into
DNADynamo and a features
map displayed. Features may
be selected and/or set as the
ORF. Multi-partite (spliced)
features may be 'joined' and
opened in a new window. The
sequence display is set to
'translated mode' and is
illustrating the start of the 'vif'
gene, together with a selected
oligo.
abi/scf chromatogram sequence alignment and editing
Editable sequence alignments in
the upper display area are linked
to chromatogram data in the lower
display area. A graphic map allows
easy visualization of sequencing
discrepancies while the 'guide'
sequence is translated and
illustrates annotations and oligos
selected from the 'features' map
automatically add vector features and draw maps
DNADynamo’s circular maps are fully
editable. You can adjust the size,
colour and position of pretty much
everything, including setting your
favourite font, as well as adding
custom text labels. DNADynamo
maps also support a special MCS
display, and allow you to add a list of
features and/or a list of absent
restriction sites.
Additional Features
•
Oligo Editor and Database
•
Integrated web BLAST search and annotation
•
Annotated multiple sequence alignment via the DNADynamo ClustalW interface
•
Pattern Searching
•
Support for demo and licensed users - we answers all your questions as rapidly as we can
more information on DNA Dynamo software features and functions is available on the Getting Started with DNA Dynamo
page, and the Analysing Sequencing Data page,
DNA Dynamo software is available for OSX, Windows and Linux. Files created on one operating system are compatible
with other operating systems, making DNA Dynamo an ideal choice for mixed operating system environments.
Copyright © BlueTractorSoftware Ltd, all rights reserved




plan sub-cloning with fragment copy and replace, or
use the dedicated ‘Construct Maker

An EcoRI/BamHI fragment containing
the vpr gene is shown selected. The
fragment can be copied and then used
to replace a selected EcoRI/BamH1
fragment in another vector
Design primers manually or use the primer designer to
automatically select primers with given properties eg for
gibson/infusion cloning or to add restriction sites
A fragment has been selected for use
in the Primer Designer - and primers
to add BamH1 sites and EcoRI sites
designed

DNA Dynamo software features
sequence navigation and restriction site
analysis
Genbank accession U52953 representing 'HIV-1 92BR025 from Brazil,
complete genome' was imported into DNADynamo and a features map
displayed. Features may be selected and/or set as the ORF. Multi-partite
(spliced) features may be 'joined' and opened in a new window. The sequence
display is set to 'translated mode' and is illustrating the start of the 'vif' gene,
together with a selected oligo.
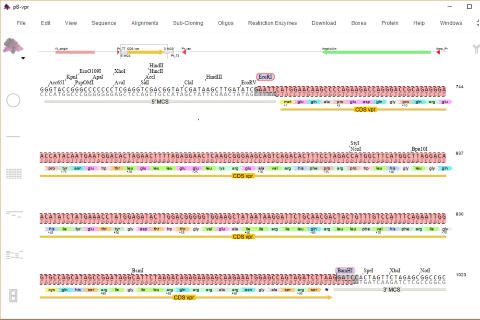
abi/scf chromatogram sequence
alignment and editing
Editable sequence alignments in the upper display area are linked to
chromatogram data in the lower display area. A graphic map allows easy
visualization of sequencing discrepancies while the 'guide' sequence is translated
and illustrates annotations and oligos selected from the 'features' map
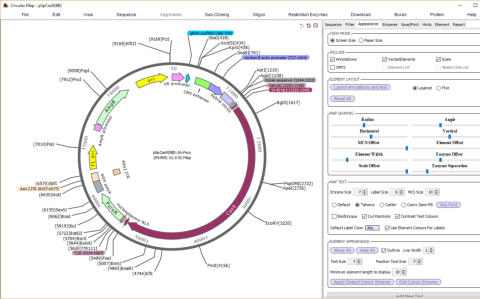
automatically add vector features and
draw maps
DNADynamo’s circular maps are fully editable. You can adjust the size, colour and
position of pretty much everything, including setting your favourite font, as well as
adding custom text labels. DNADynamo maps also support a special MCS display,
and allow you to add a list of features and/or a list of absent restriction sites.
Additional Features
•
Oligo Editor and Database
•
Integrated web BLAST search and annotation
•
Annotated multiple sequence alignment via the DNADynamo ClustalW
interface
•
Pattern Searching
•
Support for demo and licensed users - we answers all your questions as
rapidly as we can
more information on DNA Dynamo software features and functions is available on
the Getting Started with DNA Dynamo page, and the Analysing Sequencing Data
page,
DNA Dynamo software is available for OSX, Windows and Linux. Files created on
one operating system are compatible with other operating systems, making DNA
Dynamo an ideal choice for mixed operating system environments.
Copyright © BlueTractorSoftware Ltd, all rights reserved

plan sub-cloning with fragment copy and
replace, or use the dedicated ‘Construct
Maker
An EcoRI/BamHI fragment containing the vpr gene is shown selected. The
fragment can be copied and then used to replace a selected EcoRI/BamH1
fragment in another vector
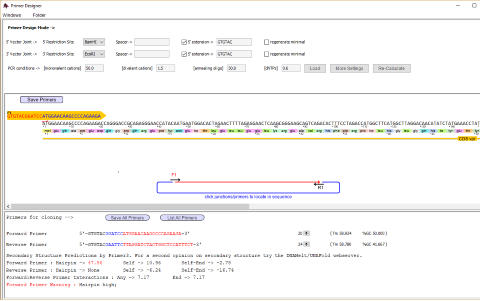
Design primers manually or use the
primer designer to automatically select
primers with given properties eg for
gibson/infusion cloning or to add
restriction sites
A fragment has been selected for use in the Primer Designer - and primers to add
BamH1 sites and EcoRI sites designed
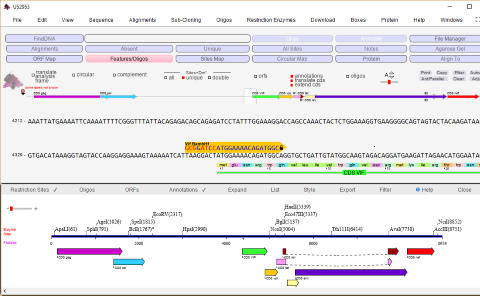
DNA Dynamo software features
sequence navigation and restriction site analysis
Genbank accession U52953 representing 'HIV-1 92BR025
from Brazil, complete genome' was imported into
DNADynamo and a features map displayed. Features may be
selected and/or set as the ORF. Multi-partite (spliced)
features may be 'joined' and opened in a new window. The
sequence display is set to 'translated mode' and is illustrating
the start of the 'vif' gene, together with a selected oligo.
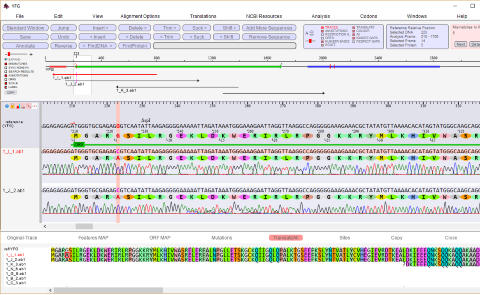
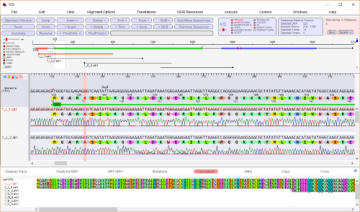
abi/scf chromatogram sequence alignment and editing
Editable sequence alignments in the upper display area are
linked to chromatogram data in the lower display area. A
graphic map allows easy visualization of sequencing
discrepancies while the 'guide' sequence is translated and
illustrates annotations and oligos selected from the 'features'
map
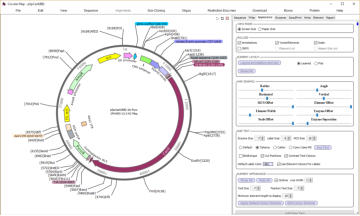
automatically add vector features and draw
maps
DNADynamo’s circular maps are fully editable. You can
adjust the size, colour and position of pretty much everything,
including setting your favourite font, as well as adding custom
text labels. DNADynamo maps also support a special MCS
display, and allow you to add a list of features and/or a list of
absent restriction sites.
Additional Features
•
Oligo Editor and Database
•
Integrated web BLAST search and annotation
•
Annotated multiple sequence alignment via the
DNADynamo ClustalW interface
•
Pattern Searching
•
Support for demo and licensed users - we answers
all your questions as rapidly as we can
more information on DNA Dynamo software features and
functions is available on the Getting Started with DNA
Dynamo page, and the Analysing Sequencing Data page,
DNA Dynamo software is available for OSX, Windows and
Linux. Files created on one operating system are compatible
with other operating systems, making DNA Dynamo an ideal
choice for mixed operating system environments.
Copyright © BlueTractorSoftware Ltd, all rights reserved

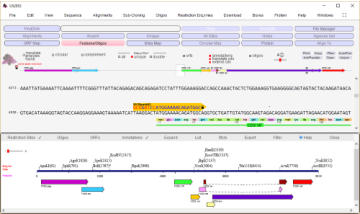
plan sub-cloning with fragment copy and replace, or
use the dedicated ‘Construct Maker
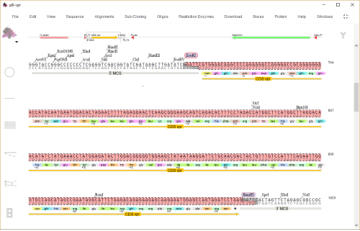
An EcoRI/BamHI fragment containing the vpr gene is shown
selected. The fragment can be copied and then used to
replace a selected EcoRI/BamH1 fragment in another vector
Design primers manually or use the primer designer to
automatically select primers with given properties eg
for gibson/infusion cloning or to add restriction sites
A fragment has been selected for use in the Primer Designer
- and primers to add BamH1 sites and EcoRI sites designed